
ABSTRACT. Chromobacterium violaceum is a free-living bacterium commonly found in aquatic habitats of tropical and subtropical regions of the world. This bacterium is able to produce a large variety of products of biotechnological and pharmacological use. Although C. violaceum is considered to be non-pathogenic, some cases of severe infections in humans and other animals have been reported. Genomic data on the type strain ATCC 12472T has provided a comprehensive basis for detailed studies of pathogenicity, virulence and drug resistance genes. A large number of open reading frames associated with various mechanisms of drug resistance were found, comprising a remarkable feature of this organism. Amongst these, beta-lactam (penicillin and cephalosporin) and multidrug resistance genes (drug efflux pumps) were the most numerous. In addition, genes associated with bacitracin, bicyclomycin, chloramphenicol, kasugamycin, and methylenomycin were also found. It is postulated that these genes contribute to the ability of C. violaceum to compete with other bacteria in the environment, and also may help to explain the common drug resistance phenotypes observed in infections caused by this bacterium. Key words: Antibiotics, Resistance, Chromobacterium violaceum INTRODUCTION Antibiotics are characterized by their chemical composition and mode of action in the target organism. The main targets for action in bacteria include the prokaryotic cell wall, the cell membrane, protein synthesis, enzymatic pathways, and DNA replication (Brooks et al., 2001). There are many different mechanisms by which microorganisms can express resistance to antibiotics, and some are very well documented: 1) they may produce enzymes that destroy the active drug, 2) they can change membrane permeability to block entry or express transport systems that actively pump the drug out into the extracellular environment, 3) they express altered structural targets, 4) they express metabolic pathways that bypass the reaction inhibited by the drug, or 5) they express altered enzymes that are less affected by the drug (Salyers and Whitt, 2001). The origin of drug resistance may be genetic or non-genetic. Active replication of bacteria is usually required for the action of most antibacterial drugs. Consequently, microorganisms that are metabolically inactive (non-multiplying or sporulated) may be phenotypically resistant to drugs. Also, microorganisms may lose the specific structural drug target for several generations, and thus become temporarily resistant. These non-genetic mechanisms of drug resistance (Brooks et al., 2001), though important in many circumstances, are temporary reversible phenotypical states. However, drug resistance that emerges as a result of genetic change and subsequent selection by antimicrobial drugs, and thus may become permanent in the bacterium and be passed onto its next generations, comprises one of the greatest problems in human and veterinary medical care (Shao et al., 2002; Perera et al., 2003), and can scale up into multidrug resistance problems (Martinez et al., 2000). Antibiotic resistance can be mediated by chromosomal and extrachromosomal genes, which can be transferred to other cells by the general mechanisms of genetic recombination, such as transduction, transformation, conjugation, and transposition (Salyers and Whitt, 2001). Some recent data suggest that different species of bacteria can develop a “mutator phenotype” under stress conditions, favoring the production of mutations associated with drug resistance (Oliver et al., 2000). In some bacteria, the expression of antibiotic production pathways, and their drug resistance counterparts, are under the control of signal sensing systems, such as quorum sensing, in which production is dependent upon the cell cycle stage, on concentrations of signals associated with high population density or on factors excreted by competing bacteria (Strauss, 1999). Chromobacterium violaceum is a Gram-negative bacillus that can be isolated from aquatic habitats in tropical and sub-tropical regions (Richard, 1993). It can sometimes cause life-threatening infections in humans and other animals, commonly sepsis, followed by cutaneous involvement and liver abscesses, though diarrheal episodes have also been reported (Liu et al., 1989; Richard, 1993; Ti et al., 1993; Chong and Lam, 1997; Midani and Rathore, 1998; Martinez et al., 2000; Dromigny et al., 2002). These are relatively rare occurrences, but some cases of septicemic diseases have been reported in South America (Petrillo et al., 1984; Kaufman et al., 1986; Martinez et al., 2000). In fact, pathogenicity islands and other virulence genes were identified in the C. violaceum genome, demonstrating that this bacterium does have a genetic potential for being pathogenic (Vasconcelos et al., 2003). Another remarkable characteristic of C. violaceum is the resistance to a relatively broad range of antibiotics (rifampin, vancomycin, ampicilin, cephalosporins), a phenomenon that makes the treatment of infections difficult (Aldridge et al., 1988; Berkowitz and Metchock, 1995; Martinez et al., 2000). The completion of genomic sequencing of C. violaceum (Vasconcelos et al., 2003) created a sound basis for studying antibiotic resistance under a new perspective, that of comparative genomics. Herein, we describe the results of a comprehensive study of C. violaceum ATCC 12472T, focusing upon genes associated with drug resistance identified in the genome. Open reading frame (ORF) similarity analyses revealed a large number of sequences in C. violaceum ATCC 12472T related to known antibiotic resistance genes. These are related to beta-lactam (Table 1), multidrug (Table 2) and other resistance genes (Table 3).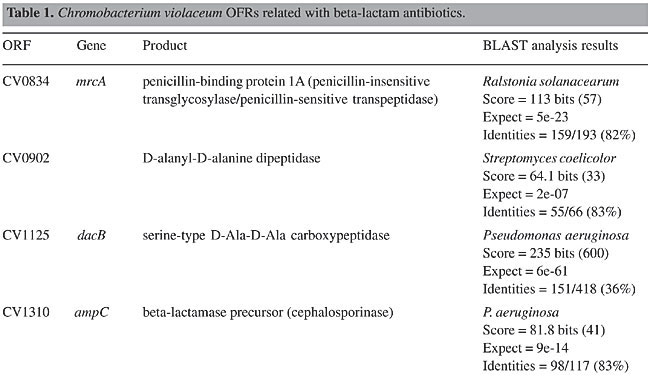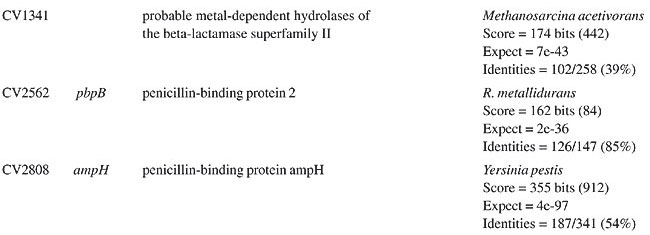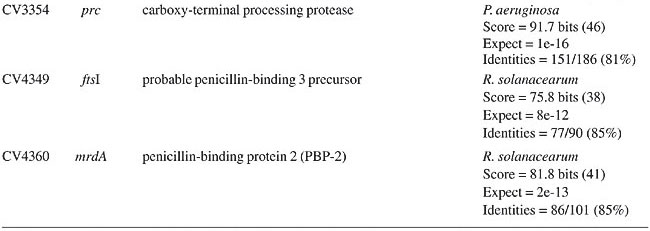    
ORF = open reading frame determined in the Chromobacterium violaceum genome (Vasconcelos et al., 2003).
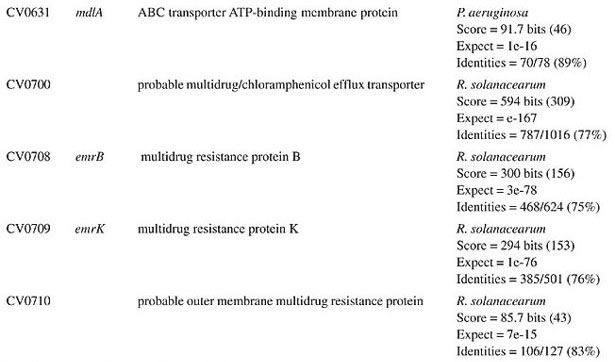
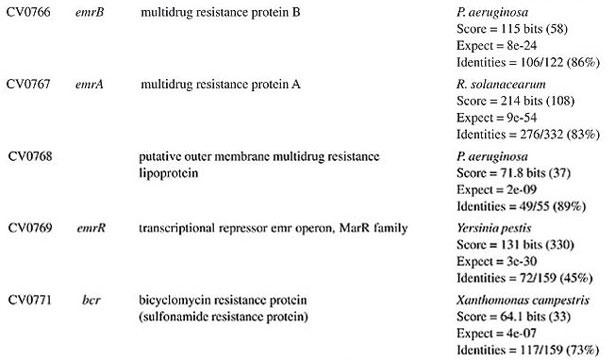
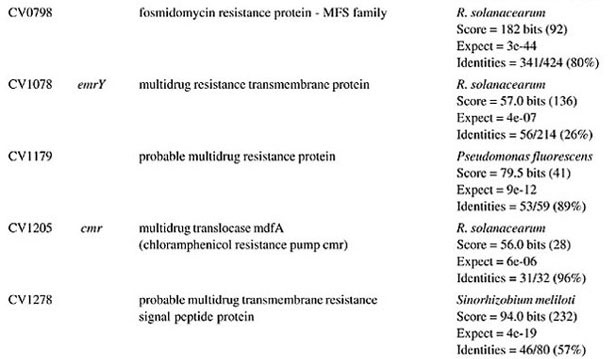
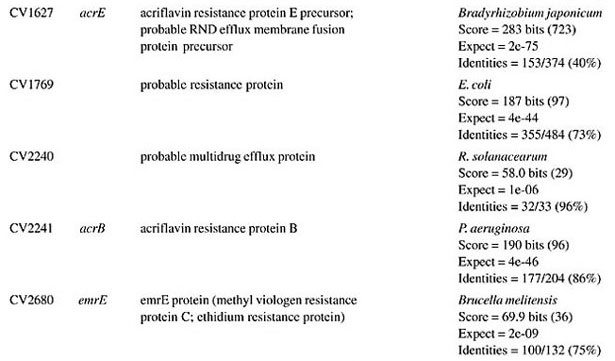
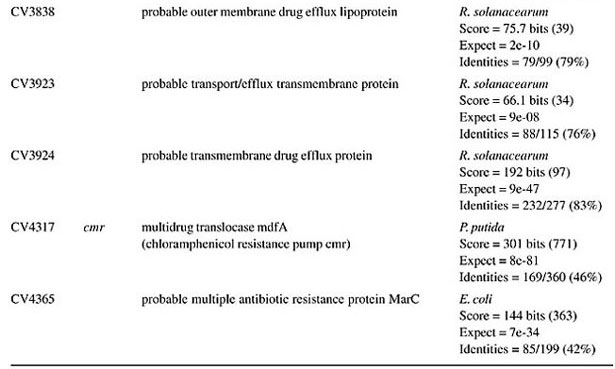
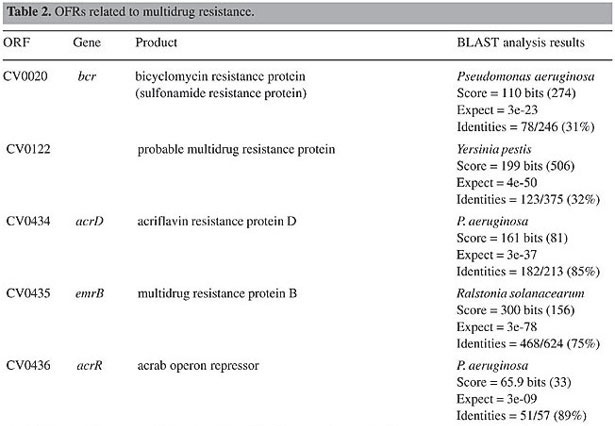       
ORF = open reading frame determined in the Chromobacterium violaceum genome (Vasconcelos et al., 2003).
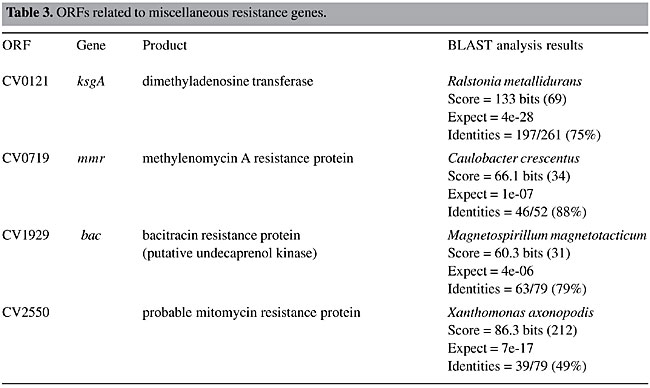
ORF = open reading frame determined in the Chromobacterium violaceum genome (Vasconcelos et al., 2003).
BETA-LACTAM ANTIBIOTIC (PENICILLIN AND CEPHALOSPORIN)- RESISTANCE GENES All beta-lactam antibiotics are selective inhibitors of bacterial cell wall synthesis by interaction with the penicillin-biding proteins. After a beta-lactam antibiotic has attached to one or more penicillin receptors, the transpeptidation reaction is inhibited and peptidoglycan synthesis is blocked. This inhibition may be due to a structural similarity of these drugs to acyl-D-alanyl-D-alanine. The next step may involve inactivation of an inhibitor of autolytic enzymes in the cell wall. Resistance to this class of antibiotics can be due to the absence of some penicillin receptors (penicillin-biding proteins), and occurs as a result of chromosomal mutations, or from failure of the beta-lactam drug to activate the autolytic enzymes. Thus, the presence of penicillin-biding proteins per se is an essential characteristic of bacteria, and does not necessarily indicate resistance. Genes that exhibit similarity to penicillin-binding proteins and to D-alanyl-D-alanine-endopeptidases of different bacterial species were identified in the C. violaceum genome (Table 1). These genes were expected, as they code for essential membrane proteins involved in the biosynthesis of murein and peptidoglycan, protein components of the cell wall. Sequence similarities indicate that one domain of these proteins belongs to a large family of beta-lactam-recognizing proteins, which includes the active-site serine beta-lactamases (Pares et al., 1996). Resistance to beta-lactams may also be determined by the organism’s production of penicillin-destroying enzymes (beta-lactamases). Beta-lactamases open the beta-lactam ring of the penicillins and cephalosporins, destroying their antimicrobial activity. There are genes with similarity to beta-lactamases or precursor proteins in the genome of C. violaceum (Table 1), suggesting that this bacterium is able to produce such proteins. D-Ala-D-Ala peptidase is a bacterial serine protease, a proteolytic enzyme with serine in its catalytic activity. These peptidases are ubiquitous, being found in viruses, bacteria and eukaryotes. They are involved in a wide range of peptidase activities, including exo-, endo-, oligo- and omega-peptidase activity. Over 20 families (denoted S1 to S27) of serine protease have been identified, being grouped into six classes (SA, SB, SC, SE, SF, and SG) on the basis of structural similarity and other functional features. Structures are known for classes SA, SB, SC, and SE. They appear to be unrelated, suggesting at least four evolutionary origins of serine peptidases, and possibly more. D-Ala-D-Ala carboxypeptidases (S11 family) are involved in the metabolism of cell components. They are synthesized with a leader peptide that targets the protein to the cell membrane. After cleavage of the leader peptide, the enzyme is retained in the membrane by a C-terminal anchor. There are three families of serine-type D-Ala-D-Ala peptidases, which are also known as low-molecular weight penicillin-binding proteins (Rawlings and Barrett, 1994). MULTIDRUG RESISTANCE SYSTEMS Drug efflux is a major mechanism of multiple drug resistance in bacteria. Generally, this is accomplished by efflux systems, responsible for unidirectional pumping of cytotoxic drugs to the extracellular environment. Although these efflux systems are usually chromosomally encoded, some may be present on plasmids. In addition to antibiotics, these pumps may export numerous dyes, detergents, disinfectants, inhibitors, organic solvents, and also homoserine lactones, involved in quorum sensing, a signaling system present in C. violaceum (Poole, 2001). Five families of drug extrusion translocases have been identified based on sequence similarity. These are the MF (major facilitator), SMR (small multidrug resistance), RND (resistance nodulation cell division), ABC (ATP-binding cassette), and the recently identified MATE (multidrug and toxic compound extrusion) family (Nikaido, 1996; Brown et al., 1999). The genome of C. violaceum contains genes with similarity to the mar (multiple antibiotic resistance) locus (Table 2). This locus is composed of the marRAB operon, which controls multiple antibiotic resistance in E. coli. MarR (slyA) is a regulator, from the XylS/AraC family of transcriptional regulator proteins, whose product (MarR protein) alters the expression of several target genes. Structural studies showed MarR as a dimmer, with each subunit containing a winged-helix DNA binding motif (Alekshun et al., 2001). Numerous bacterial transcription regulatory proteins bind DNA via a helix-turn-helix motif. These proteins are very diverse, but for convenience may be grouped into subfamilies on the basis of sequence similarity. One such family, marR is grouped with a wide range of proteins, including emrR, hpcR, hpR, marR, pecS, petP, papX, prsX, ywaE, yxaD, and yybA. The Mar proteins are involved in multiple antibiotic resistance, a non-specific resistance system. The expression of the mar operon is controlled by a repressor, MarR, which interacts with a large number of compounds in order to induce transcription of the mar operon, possibly via loss of its DNA binding ability. C. violaceum also contains an ORF with similarity to marC. Although the function of this gene is unknown, similarity with efflux protein genes of E. coli suggests the same activity for it. Genes with similarity to the emrRAB locus of E. coli are also present in C. violaceum (Lomovskaya and Lewis, 1992). This operon encodes a multidrug resistance pump of the MF-family that protects the cell from several chemically unrelated antimicrobial agents, e.g., the protonophores carbonyl cyanide m-chlorophenylhydrazone (CCCP), tetrachlorosalicyl anilide and the antibiotics, nalidixic acid and thiolactomycin. The emrR is the first gene of the operon, and was shown to be a repressor of microcin biosynthesis (del Castillo et al., 1990). Overproduction of the EmrR protein (with a multicopy vector containing the cloned emrR gene) suppressed transcription of the emr operon. A mutation in the emrR gene led to over-expression of the EmrAB pump and increased resistance to antimicrobial agents. CCCP, nalidixic acid, and a number of other structurally unrelated chemicals induced expression of the emr genes, and the induction required an EmrR regulator. In conclusion, the emrRAB genes constitute an operon, and EmrR serves as its negative regulator. Some of the chemicals that induce the pump serve as its substrates, suggesting that their extrusion is the natural function of the pump (Lomovskaya et al., 1995). Proteins of the SMR family are unusually small and are predicted to span the membrane four times. The emrE (mvrC) gene, the disruption of which causes hyper-susceptibility to tetraphenylphosphonium, methylviologen, and ethidium bromide, is a member of this family in E. coli. Overproduction of EmrE protein makes E. coli slightly more resistant to tetracycline, erythromycin, and sulfadiazine (Nikaido, 1996). C. violaceum contains a gene similar to emrE. Some genes of C. violaceum exhibit similarity to the adenosine triphosphate (ATP)-binding transport system (ABC transporters), such as mdlA and B (Table 2). These proteins contain an (ATP)-biding cassette (ABC), which is also called a nucleotide-biding domain. They are all integral membrane proteins involved in a variety of transport systems that require ATP. Members of this family include the cystic fibrosis transmembrane conductance regulator, bacterial leukotoxin secretion ATP-binding protein, multidrug resistance proteins, the yeast leptomycin B resistance protein, the mammalian sulfonylurea receptor, and antigen peptide transporter 2 (van Veen et al., 1996). The genes ylcB, acrR, acrA, and acrB (Table 2) exhibit similarity to the MexAB-OprM broadly-specific multidrug efflux system of Pseudomonas aeruginosa and E. coli (Ma et al., 1996; Poole, 2001; Nikaido and Zgurskaya, 2001). These proteins belong to the RND family and mediate resistance to different drugs, including acriflavine, a macrolide that has homology with flavokinase. In E. coli, the acrA and acrB genes encode a multidrug efflux system that is believed to protect the bacterium against hydrophobic inhibitors. AcrR is a regulator of acrAB genes, acting as a modulator to fine tune the level of acrAB transcription. This type of gene regulation is uncommon in E. coli. The acrAB expression is up-regulated under general stress conditions, but this activation is not likely to be directly mediated by the known global stress regulators, such as MarA or SoxS, although elevated levels of these proteins were shown to increase the transcription of acrAB (Ma et al., 1996). CHLORAMPHENICOL MdfA, the product of the mdfA gene homolog identified in the C. violaceum genome (Table 2), is an MF-related protein, the over-expression of which results in resistance to a diverse group of cationic and zwitterionic lipophilic compounds, such as ethidium bromide, puromycin, rifampicin, tetracycline, tetraphenylphosphonium, and also to chemically unrelated, clinically important antibiotics, such as chloramphenicol and erythromycin. Some studies have proposed that MdfA is a drug proton antiporter (Edgar and Bibi, 1999). BICYCLOMYCIN The bicyclomycin resistance protein (Bcr) is a multidrug transporter from the MF family. The growth-inhibiting drug bicyclomycin is known to be an inhibitor of Rho factor activity in E. coli (Yanofsky and Horn, 1995). The C. violaceum bcr gene presents similarity to the corresponding gene of Pseudomonas aeruginosa (Table 2). MISCELLANEOUS RESISTANCE GENES Bacitracin Bacitracin resistance protein (BacA) is a putative undecaprenol kinase that confers resistance to bacitracin, probably by phosphorylation of undecaprenol (Cain et al., 1993). The bacA gene of C. violaceum presents similarity to the bacA gene of Magnetospirillum magnetotacticum (Table 3). Kasugamycin The 16S rRNA dimethylase (gene ksgA) acts in the biogenesis of ribosomes by catalyzing the dimethylation of two adjacent adenosines in the loop of a conserved hairpin near the 3'-end of the 16S rRNA. Inactivation of ksgA leads to resistance to the aminoglycoside antibiotic kasugamycin. The gene found in C. violaceum is similar to the Ralstonia metallidurans ksgA gene (Table 3) (Vila-Sanjurjo et al., 1999). Methylenomycin ORF CV0719 of the C. violaceum genome shows homology with the methylenomycin resistance protein (MMR) peptide (Table 3). The predicted mmr-specified protein should be hydrophobic and should present homology at the amino acid level to tetracycline-resistance proteins from both Gram-positive and Gram-negative bacteria. It is suggested that methylenomycin resistance may be conferred by a membrane protein, perhaps controlling the efflux of the antibiotic (Neal and Chater, 1987). CONCLUDING REMARKS A remarkable characteristic of C. violaceum is the wide range of multidrug resistance and other antibiotic resistance gene homologues in its genome. Secretion of different compounds is a postulated feature of this organism. Certainly, these systems play important roles in the physiology of the bacterium. Interestingly, as these pumps may also transport homoserine lactones, they could have additional roles in cell signaling, such as quorum sensing, a property already demonstrated for C. violaceum. The presence of penicillin-binding proteins in C. violaceum is an expected characteristic, because these proteins play an essential role in the cell wall biosynthesis. On the other hand, the presence of beta-lactamase precursors could indicate resistance to beta-lactam antibiotics. This is an important characteristic that probably improves the capacity of C. violaceum to compete with other microorganisms in the environment. Additionally, we cannot rule out a function of the beta-lactamases in the rare cases of infections in humans. The presence of genes that mediate resistance to several other antibiotics was deduced by similarity with counterparts in other bacteria, indicating that drug resistance is an important characteristic of C. violaceum. ACKNOWLEDGMENTS F. Fantinatti-Garboggini, P.B. Trevilato, R. Almeida, T.A. P. Barbosa and V.A. Portillo, were supported by grants from the CNPq/RHAE. The Chromobacterium violaceum genome sequencing project was supported by funds from the MCT (Ministério da Ciência e Tecnologia) and CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) in the Brazilian National Genome Project Consortium. REFERENCES Aldridge, K.E., Valainis, G.T. and Sanders, C.V. (1988). Comparison of the in vitro activity of ciprofloxacin and 24 other antimicrobial agents against clinical strains of Chromobacterium violaceum. Diagn. Microbiol. Infect. Dis. 10: 31-39. Alekshun, M.N., Levy, S.B., Mealy, T.R., Seaton, B.A. and Head, J.F. (2001). The crystal structure of MarR, a regulator of multiple antibiotic resistance, at 2.3 A resolution. Nat. Struct. Biol. 8: 710-714. Berkowitz, F.E. and Metchock, B. (1995). Third generation cephalosporin-resistant gram-negative bacilli in the feces of hospitalized children. Pediatr. Infect. Dis. J. 14: 97-100. Brooks, G.F., Butel, J.S. and Morse, S.A. (2001). Medical Microbiology. McGraw-Hill/Appleton & Lange, New York, NY, USA. Brown, M.H., Paulsen, I.T. and Skurray, R.A. (1999). The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol. Microbiol. 31: 394-395. Cain, B.D., Norton, P.J., Eubanks, W., Nick, H.S. and Allen, C.M. (1993). Amplification of the bacA gene confers bacitracin resistance to Escherichia coli. J. Bacteriol. 175: 3784-3789. Chong, C.Y. and Lam, M.S. (1997). Case report and review of Chromobacterium sepsis - a gram-negative sepsis mimicking melioidosis. Singapore Med. J. 38: 263-265. del Castillo, I., Gomez, J.M. and Moreno, F. (1990). mprA, an Escherichia coli gene that reduces growth-phase-dependent synthesis of microcins B17 and C7 and blocks osmoinduction of proU when cloned on a high-copy-number plasmid. J. Bacteriol. 172: 437-445. Dromigny, J.A., Fall, A.L., Diouf, S. and Perrier-Gros-Claude, J.D. (2002). Chromobacterium violaceum: a case of diarrhea in Senegal. Pediatr. Infect. Dis. J. 21: 573-574. Edgar, R. and Bibi, E. (1999). A single membrane-embedded negative charge is critical for recognizing positively charged drugs by the Escherichia coli multidrug resistance protein MdfA. EMBO J. 18: 822-832. Kaufman, S.C., Ceraso, D. and Shugurensky, A. (1986). First case report from Argentina of a fatal septicemia caused by Chromobacterium violaceum. J. Clin. Microbiol. 23: 956-958. Liu, C.H., Chu, R.M., Weng, C.N., Lin, Y.L. and Chi, C.S. (1989). An acute pleuropneumonia in a pig caused by Chromobacterium violaceum. J. Comp. Pathol. 100: 459-463. Lomovskaya, O. and Lewis, K. (1992). Emr, an Escherichia coli locus for multidrug resistance. Proc. Natl. Acad. Sci. USA 89: 8938-8942. Lomovskaya, O., Lewis, K. and Matin, A. (1995). EmrR is a negative regulator of the Escherichia coli multidrug resistance pump EmrAB. J. Bacteriol. 177: 2328-2334. Ma, D., Alberti, M., Lynch, C., Nikaido, H. and Hearst, J.E. (1996). The local repressor AcrR plays a modulating role in the regulation of acrAB genes of Escherichia coli by global stress signals. Mol. Microbiol. 19: 101-112. Martinez, R., Velludo, M.A., Santos, V.R. and Dinamarco, P.V. (2000). Chromobacterium violaceum infection in Brazil. A case report. Rev. Inst. Med. Trop. São Paulo 42: 111-113. Midani, S. and Rathore, M. (1998). Chromobacterium violaceum infection. South. Med. J. 91: 464-466. Neal, R.J. and Chater, K.F. (1987). Nucleotide sequence analysis reveals similarities between proteins determining methylenomycin A resistance in Streptomyces and tetracycline resistance in eubacteria. Gene 58: 229-241. Nikaido, H. (1996). Multidrug efflux pumps of gram-negative bacteria. J. Bacteriol. 178: 5853-5859. Nikaido, H. and Zgurskaya, H.I. (2001). AcrAB and related multidrug efflux pumps of Escherichia coli. J. Mol. Microbiol. Biotechnol. 3: 215-218. Oliver, A., Cantón, R., Campo, P., Baquero, F. and Blázquez, J. (2000). High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science 288: 1251-1253. Pares, S., Mouz, N., Petillot, Y., Hakenbeck, R. and Dideberg, O. (1996). X-ray structure of Streptococcus pneumoniae PBP2x, a primary penicillin target enzyme. Nat. Struct. Biol. 3: 218-219. Perera, S., Punchihewa, P.M., Karunanayake, M.C. and de Silva, N. (2003). Fatal septicaemia caused by Chromobacterium violaceum. Ceylon Med J. 48: 26-27. Petrillo, V.F., Severo, V., Santos, M.M. and Edelweiss, E.L. (1984). Recurrent infection with Chromobacterium violaceum: first case report from South America. J. Infect. 9: 167-169. Poole, K. (2001). Multidrug efflux pumps and antimicrobial resistance in Pseudomonas aeruginosa and related organisms. J. Mol. Microbiol. Biotechnol. 3: 255-264. Rawlings, N.D. and Barrett, A.J. (1994). Families of serine peptidases. Methods. Enzymol. 244: 19-61. Richard, C. (1993). Chromobacterium violaceum, opportunist pathogenic bacteria in tropical and subtropical regions. Bull. Soc. Pathol. Exot. 86: 169-173. Salyers, A.A. and Whitt, D.D. (2001). Bacterial Pathogenesis: A Molecular Approach. 2nd edn. ASM Press, Washington, D.C., USA. Shao, P.L., Hsueh, P.R., Chang, Y.C., Lu, C.Y., Lee, P.Y., Lee, C.Y. and Huang, L.M. (2002). Chromobacterium violaceum infection in children: a case of fatal septicemia with nasopharyngeal abscess and literature review. Pediatr. Infect. Dis. J. 21: 707-709. Strauss, E. (1999). A symphony of bacterial voices. Science 284: 1302-1304. Ti, T.Y., Tan, W.C., Chong, A.P. and Lee, E.H. (1993). Nonfatal and fatal infections caused by Chromobacterium violaceum. Clin. Infect. Dis. 17: 505-507. van Veen, H.W., Venema, K., Bolhuis, H., Oussenko, I., Kok, J., Poolman, B.A.J. and Konings, W.N. (1996). Multidrug resistance mediated by a bacterial homologue of the human multidrug transporter MDR1. Proc. Natl. Acad. Sci. USA 93: 10668-10672. Vasconcelos, A.T.R., Almeida, D.F., Almeida, F.C., Almeida, L.G.P., Almeida, R., Alves-Gomes, J.A., Andrade, E.M., Antônio, R.V., Araripe, J., Araújo, M.F.F., Astolfi-Filho, S., Azevedo, V., Baptista, A.J., Bataus, L.A.M., Batista, J.S., Beló, A., van den Berg, C., Blamey, J., Bogo, M., Bonatto, S., Bordignon, J., Brito, C.A., Brocchi, M., Burity, H.A., Camargo, A.A., Cardoso, D.D.P., Carneiro, N.P., Carraro, D.M., Carvalho, C.M.B., Cascardo, J.C.M., Cavada, B.S., Chueire, L.M.O., Creczynski-Pasa, T.B., Duran, N., Fagundes, N., Falcão, C.L., Fantinatti, F., Farias, I.P., Felipe, M.S.S., Ferrari, L.P., Ferro, J.A., Ferro, M.I.T., Franco, G.R., Freitas, N.S.A., Furlan, L.R., Gazzinelli, R.T., Gomes, E.A., Gonçalves, P.R., Grangeiro, T.B., Grattapaglia, D., Grisard, E.C., Guimarães, C.T., Hanna, E.S., Hungria, M., Jardim, S.N., Laurino, J., Leoi, L.C.T., Lima, L.F.A., Loureiro, M.F., Lyra, M.C.C.P., Macedo, M., Madeira, H.M.F., Manfio, G.P., Maranhão, A.Q., Martins, W.S., di Mauro, S.M.Z., Medeiros, S.R.B., Meissner, R.V., Moreira, M.A.M., Nascimento, F.F., Nicolas, M.F., Oliveira, J.G., Oliveira, S.C., Paixão, R.F.C., Parente, J.A., Pedrosa, F.O., Pena, S.D.J., Pereira, J.O., Pereira, M., Pinto, L.S.R.C., Pinto, L.S., Porto, J.I.R., Potrich, D.P., Ramalho-Neto, C.E., Reis, A.M.M., Rigo, L.U., Rondinelli, E., Santos, E.B.P., Santos, F.R., Schneider, M.P.C., Seuanez, H.N., Silva, A.M.R., Silva, A.L.C., Silva, D.W., Silva, R., Simões, I.D.C., Simon, D., Soares, C.M.A., Soares, R.B.A., Souza, E.M., Souza, K.R.L., Souza, R.C., Steffens, M.B.R., Steindel, M., Teixeira, S.R., Urmenyi, T., Vettore, A., Wassem, R., Zaha, A. and Simpson, A.J.G. (2003). Complete genome sequence of Chromobacterium violaceum reveals remarkable and exploitable bacterial adaptability. Proc. Natl. Acad. Sci. USA 100: 11660-11665. Vila-Sanjurjo, A., Squires, C.L. and Dahlberg, A.E. (1999). Isolation of kasugamycin resistant mutants in the 16 S ribosomal RNA of Escherichia coli. J. Mol. Biol. 293: 1-8. Yanofsky, C. and Horn, V. (1995). Bicyclomycin sensitivity and resistance affect Rho factor-mediated transcription termination in the tna operon of Escherichia coli. J. Bacteriol. 177: 4451-4456. |
|